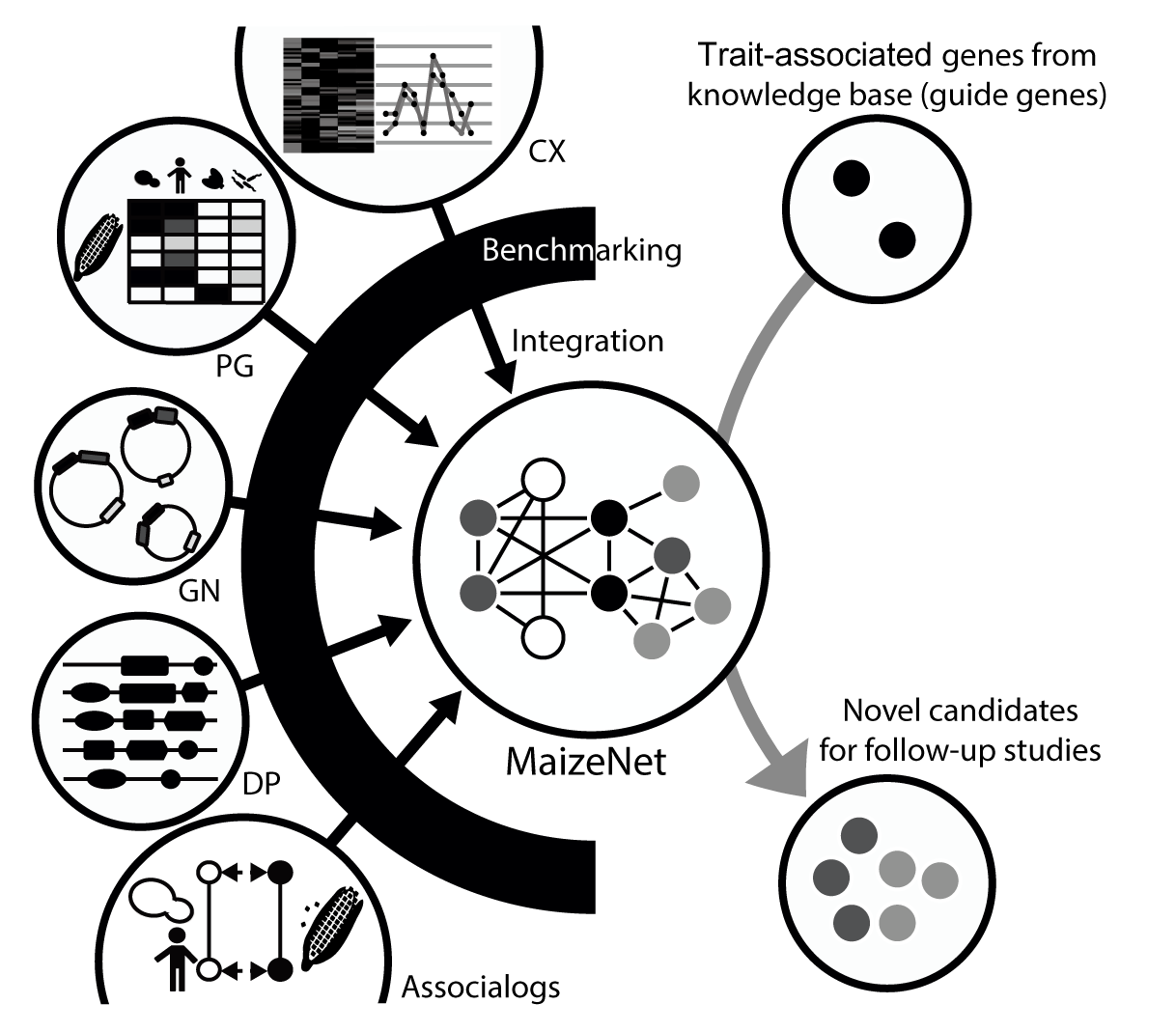
A popular strategy for facilitating genetic dissection of complex traits is prioritizing genes by network connections to known trait-associated genes that can guide network searches for novel trait-associated genes.
New candidates are then ranked by closeness to the "guide genes" measured for each candidate gene as the sum of network edge scores (Log likelihood scores) from that gene to the guide genes.
As both high-throughput forward and reverse genetics pose challenges due to the large maize genome, in silico selection of highly probable candidate genes followed by experimental validation may accelerate the mapping of gene-to-trait associations.
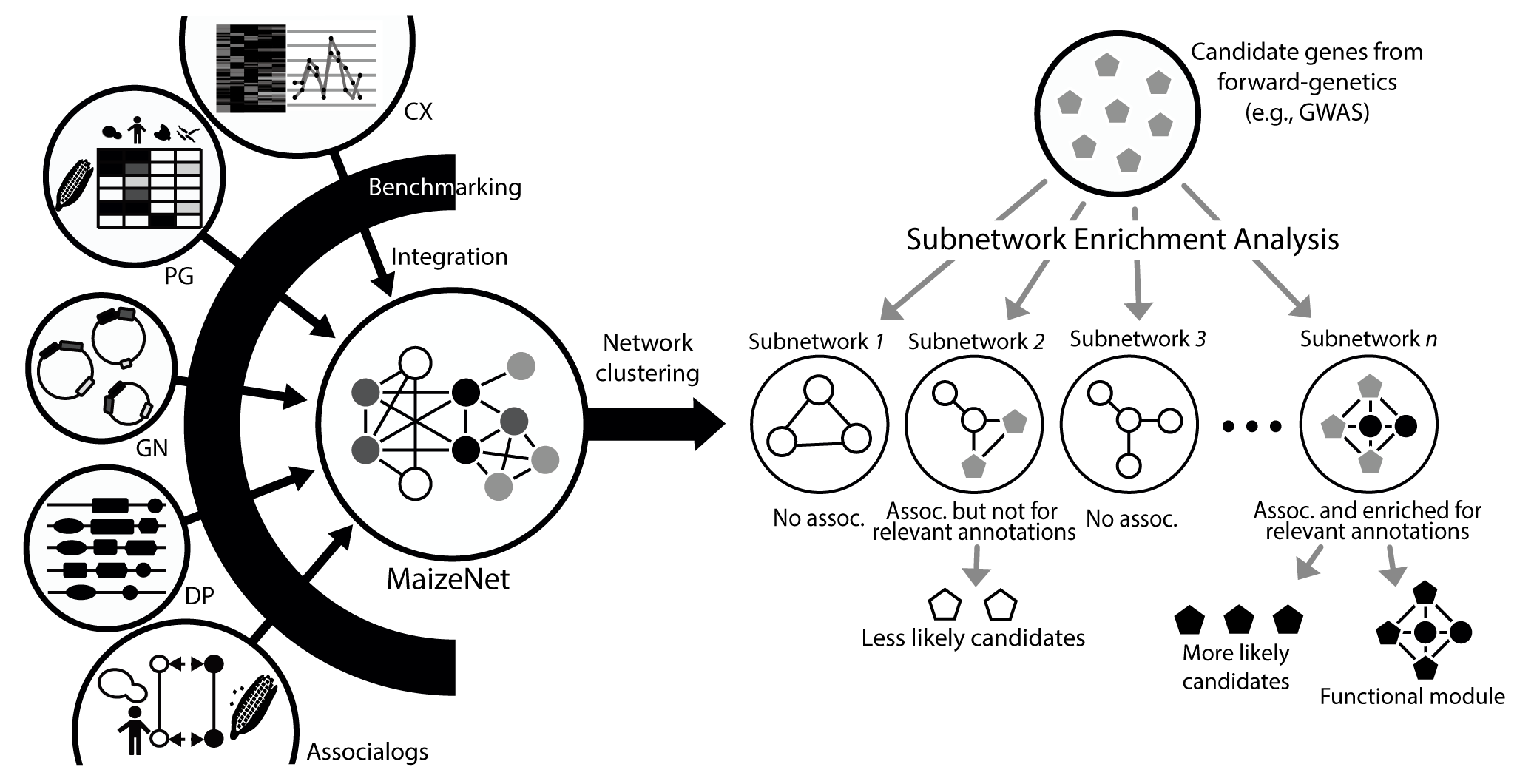
MaizeNet can stratify candidate genes from unbiased forward-genetics such as GWAS via associations with subnetworks enriched for relevant GOBP annotations.
For this purpose, we generated subnetworks of MaizeNet through overlapping Markov clustering (Shih and Parthasarathy, 2012), and measured enrichment of GWAS candidate genes across 2335 MaizeNet subnetworks containing at least 5 genes.
GWAS candidates generally contain many false positives. If significantly associated subnetworks with the given GWAS candidate genes (# overlap genes ≥ 2 and P < 0.01) are also enriched for relevant GOBP annotations, the subnetwork is likely to be a functional module for the trait and the associated candidate genes are more likely to be involved in the trait.
