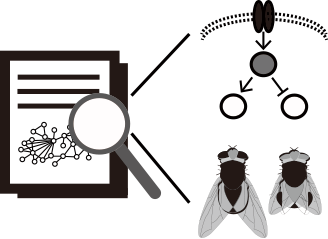

Get a functional insight for an uncharacterized Drosophila gene by looking functional annotations for network neighbors. This option collects all GO biological process and RNAi phenotype annotations among network neighbors of a query gene and lists them by the order of enrichment.
- Inferences of prioritized biological function for queried genes
- Inferences of prioritized RNAi phenotypes that associated with queried genes
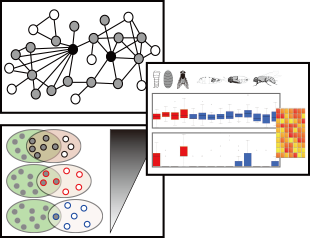
Identify new candidate genes for the function/pathway/phenotype of Drosophila by submitting a set of known genes as input data. These submitted genes are used as guide genes for gene prioritization. This option collects closely connected genes to the guide genes on the network and list the prioritized genes by total edge weight score (sum of log likelihood score) to the guide gene set.
- Estimated prediction power of guide genes within the network (AUC)
- Distribution of spatiotemporal state-specific network links
- Network graph of guide and candidate genes (Direct neighborhood only)
- Prioritized list of guide and candidate genes and those network evidences
- Gene set analysis of various functional annotations

This option is similar to Gene prioritization option, except prioritizing genes for human disease. As guide genes, both fly genes (Drosophila orthologs of human disease genes) and human disease genes are acceptable. Users can use additional option of support genes to further filtration for more confident candidates. Support genes from human genome-wide association studies and de novo mutation screens are currently available in the FlyNet server, and users can submit their own list of support genes. FlyNet reports Drosophila orthologs of the new disease genes and their cohesiveness in the network. These results assess the suitability of Drosophila for human disease models and suggest candidate Drosophila genes and RNAi phenotypes for genetic analysis of the human diseases in fly models.
- "Gene prioritization" network analysis of human genes by those Drosophila orthologs
- Prioritized candidate Drosophila RNAi phenotypes and the list of associated genes
